假单胞菌属
假单胞菌属(学名:Pseudomonas)是一类需氧的革兰氏阴性细菌,它位于假单胞菌科下,已知物种有191个。该属微生物具有极其丰富的代谢多样性,这些多样性也使得它们能够在非常广阔的生态位中生存。[1]假单胞菌属的微生物在in vitro条件下很容易培养,这使得该属成为科研的绝佳材料,比较典型的研究对象有人类致病菌绿脓杆菌、植物致病菌丁香假单胞菌、土壤细菌恋臭假单胞菌以及能促进植物生长的荧光假单胞菌。
| 假单胞菌属 | |
|---|---|
| |
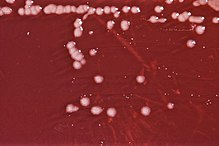
| 琼脂板上的绿脓杆菌菌落 | |
| 科学分类 | |
| 域: | 细菌域 Bacteria |
| 界: | 细菌界 Bacteria |
| 门: | 變形菌門 Pseudomonadota |
| 纲: | γ-變形菌綱 Gammaproteobacteria |
| 目: | 假單胞菌目 Pseudomonadales |
| 科: | 假单胞菌科 Pseudomonadaceae |
| 属: | 假单胞菌属 Pseudomonas Migula, 1894 |
| 模式種 | |
| 绿脓杆菌 Pseudomonas aeruginosa (Schroeter, 1872) Migula, 1900
| |
| 种 | |
|
见正文 | |
| 異名 | |
| |
因为广泛存在于水体和诸如双子叶植物等的种子中,假单胞菌属算是较早发现的一种微生物。假单胞菌属这个名字最早在1894和1900年由Walter Migula给出,当时只是一个很模糊的名词,它的意思是指“革兰氏阴性菌,杆状以及一端有鞭毛的、能够形成芽孢的细菌”。[2][3]但是后来这些被认为芽孢的东西被证明只是某些储存物质的颗粒而已。[4]现在我们已经不用这些比较模糊的描述了,而选用绿脓杆菌作为模式种来研究。[4]
分类历史 编辑
和大部分细菌属一样,假单胞菌属的最后共同祖先生活于大约几亿年前。它最早在19世纪末由Walter Migula定名,但它名字的来源在当时并未给出,而是最早见于第七版的细菌命名法著作《伯杰氏系统细菌学手册》(Bergey's Manual of Systematic Microbacteriology):pseudo来源于希腊语的pseudes(ψευδής),即“假”之义;-monas来源于μονάς/μονάδος,为“单个单元”之义;合起来可以把这个名字理解为“假的单元”之义。但是这个名字没有任何有价值的含义,它并不是说这种生物可能错误地会出现单个细胞的状态,因为它从来就没有以多细胞的形式存在过。一种比较可靠的猜测是,Migula这样起名可能单纯是想说这种生物是假的“Monas”,Monas是指金藻纲的一种有着微小鞭毛的原生生物。[4]不久之后,其他符合Migula这一模糊定义的微生物也陆续被发现并被加入到这一属下,但是它们当中的很多在后来通过检测保守性大分子等方法又被重新划分到其他门类下了。[5]
最近,利用16S rRNA测序分析手段,很多细菌都被重新划分分类。[6]有一些原来属于单胞菌属和黄色单胞菌属的细菌又被划分到了假单胞菌属下。[7]原来属于假单胞菌属的一些菌株现在被分到了伯克氏菌属和罗尔斯通氏菌属。[8][9]
2000年,人们获得了假单胞菌属某物种的全基因组序列,其后很多其他菌株的序列也被测出,包括:P. aeruginosa strains PAO1 (2000), P. putida KT2440 (2002), P. protegens Pf-5 (2005), P. syringae pathovar tomato DC3000 (2003), P. syringae pathovar syringae B728a (2005), P. syringae pathovar phaseolica 1448A (2005), P. fluorescens Pf0-1, and P. entomophila L48。[5]
下属物种 编辑
本属包括以下物种:
- 绿脓杆菌 Pseudomonas aeruginosa 组
- 绿脓杆菌 Pseudomonas aeruginosa
- 产碱假单胞菌 Pseudomonas alcaligenes
- 鳗败血假单胞菌 Pseudomonas anguilliseptica
- 阿根廷假单胞菌 Pseudomonas argentinensis
- 烂泥假单胞菌 Pseudomonas borbori
- 香茅醇假单胞菌 Pseudomonas citronellolis
- 变黄假单胞菌 Pseudomonas flavescens
- 门多萨假单胞菌 Pseudomonas mendocina
- 硝基还原假单胞菌 Pseudomonas nitroreducens
- 食油假单胞菌 Pseudomonas oleovorans
- 类产碱假单胞菌 Pseudomonas pseudoalcaligenes
- 食树脂假单胞菌 Pseudomonas resinovorans
- 稻草假单胞菌 Pseudomonas straminea
- 绿叶假单胞菌Pseudomonas chlororaphis 组
- 铁角蕨假单胞菌 Pseudomonas asplenii
- 桔黄假单胞菌 Pseudomonas aurantiaca
- 致黄假单胞菌 Pseudomonas aureofaciens
- 绿叶假单胞菌 Pseudomonas chlororaphis
- 皱纹假单胞菌 Pseudomonas corrugata
- 莓实假单胞菌 Pseudomonas fragi
- 隆德假单胞菌 Pseudomonas lundensis
- 腐臭假单胞菌 Pseudomonas taetrolens
- 荧光假单胞菌 Pseudomonas fluorescens 组
- 南极假单胞菌 Pseudomonas antarctica
- 产氮假单胞菌 Pseudomonas azotoformans
- Pseudomonas blatchfordae
- 油菜假单胞菌 Pseudomonas brassicacearum
- 布氏假单胞菌 Pseudomonas brenneri
- Pseudomonas cedrina
- 皱纹假单胞菌 Pseudomonas corrugata
- 荧光假单胞菌 Pseudomonas fluorescens
- 盖氏假单胞菌 Pseudomonas gessardii
- 黎巴嫩假单胞菌 Pseudomonas libanensis
- 孟氏假单胞菌 Pseudomonas mandelii
- 边缘假单胞菌 Pseudomonas marginalis
- 地中海假单胞菌 Pseudomonas mediterranea
- 子午线假单胞菌 Pseudomonas meridiana
- 米氏假单胞菌 Pseudomonas migulae
- 霉味假单胞菌 Pseudomonas mucidolens
- Pseudomonas orientalis
- Pseudomonas panacis
- Pseudomonas protegens
- 解朊假单胞菌 Pseudomonas proteolytica
- 罗氏假单胞菌 Pseudomonas rhodesiae
- 类黄假单胞菌 Pseudomonas synxantha
- 赛维瓦尔假单胞菌 Pseudomonas thivervalensis
- 托拉斯假单胞菌 Pseudomonas tolaasii
- 斡氏假单胞菌 Pseudomonas veronii
- 穿孔素假单胞菌Pseudomonas pertucinogena 组
- 恋臭假单胞菌Pseudomonas putida 组
- 乳脂色假单胞菌 Pseudomonas cremoricolorata
- Pseudomonas entomophila
- 黄褐假单胞菌 Pseudomonas fulva
- 蒙氏假单胞菌 Pseudomonas monteilii
- 摩氏假单胞菌 Pseudomonas mosselii
- 栖麦假单胞菌 Pseudomonas oryzihabitans
- 副黄假单胞菌 Pseudomonas parafulva
- 香鱼假单胞菌 Pseudomonas plecoglossicida
- 恋臭假单胞菌 Pseudomonas putida
- 施氏假单胞菌Pseudomonas stutzeri 组
- 丁香假单胞菌Pseudomonas syringae组
- Pseudomonas amygdali Psallidas and Panagopoulos 1975
- Pseudomonas avellanae Janse et al. 1997
- Pseudomonas caricapapayae Robbs 1956
- 菊苣假单胞菌 Pseudomonas cichorii
- Pseudomonas coronafaciens
- Pseudomonas ficuserectae Goto 1983
- Pseudomonas helianthi
- Pseudomonas meliae Ogimi, 1981,为Pseudomonas_Q kirkiae的异名
- Pseudomonas savastanoi
- 菜豆晕疫病菌 Pseudomonas savastanoi pv. phaseolicola ( Burkholder ) Gardan et al.
- 核果树溃疡病菌 Pseudomonas syringae pv. morsprunorum ( Wormald ) Young et al.
- 桃树溃疡病菌 Pseudomonas syringae pv. persicae ( Prunier et al. ) Young et al.
- 豌豆细菌性疫病菌 Pseudomonas syringae pv. pisi ( Sackett ) Young et al.
- 十字花科黑斑病菌 Pseudomonas syringae pv. maculicola (McCulloch) Young et al
- 番茄细菌性叶斑病菌 Pseudomonas syringae pv. tomato ( Okabe ) Young et al.
- 丁香假单胞菌 Pseudomonas syringae
- 西红柿假单胞菌 Pseudomonas tomato
- 浅绿黄假单胞菌 Pseudomonas viridiflava
- 组的地位未定的菌种:
- Pseudomonas abietaniphila Mohn et al., 1999
- Pseudomonas acidophila
- Pseudomonas aestuarii Kim et al. 2023[10]
- 伞菌假单胞菌 Pseudomonas agarici Young, 1970
- Pseudomonas akapageensis
- 嗜碱假单胞菌 Pseudomonas alcaliphila Yumoto et al., 2001
- Pseudomonas alkanolytica Chocat et al. 1983
- Pseudomonas amyloderamosa Norrman and Wober 1975
- Pseudomonas aphyarum Testerman et al. 2023[11]
- Pseudomonas aromaticivorans Banerjee et al. 2023[12]
- 铁角蕨假单胞菌 Pseudomonas asplenii (Ark & Tompkins, 1946) Savulescu, 1947
- Pseudomonas atagonensis Morimoto et al. 2020[13]
- Pseudomonas azotifigens Hatayama et al., 2005
- 博岑假单胞菌 Pseudomonas bauzanensis [14]
- 食苯假单胞菌 Pseudomonas benzenivorans Lang et al., 2012
- Pseudomonas benzopyrenica Dong et al. 2023[15]
- Pseudomonas bharatica Mohapatra and Phale 2023[16]
- Pseudomonas cannabina (ex Šutič and Dowson 1959) Gardan et al. 1999
- Pseudomonas coenobios
- 结冰假单胞菌 Pseudomonas congelans Behrendt et al. 2003
- 康氏假单胞菌 Pseudomonas costantinii Munsch et al. 2002
- Pseudomonas cruciviae
- Pseudomonas cucumis Liao et al. 2023[17]
- 多瑙河假单胞菌 Pseudomonas danubii Mulet et al. 2023[18]
- Pseudomonas delhiensis
- Pseudomonas excibis
- Pseudomonas extremorientalis
- Pseudomonas fontis Testerman et al. 2023[19]
- 弗雷德里克斯堡假单胞菌 Pseudomonas frederiksbergensis
- Pseudomonas fuscovaginae (ex Tanii et al. 1976) Miyajima et al. 1983
- Pseudomonas gelidicola Kadota 1951
- Pseudomonas graminis Behrendt et al. 1999[20]
- 格氏假单胞菌 Pseudomonas grimontii Baïda et al. 2002,以法国细菌学家格里蒙特(P. A. D. Grimon)的名字命名
- Pseudomonas guryensis Kim et al. 2021[21]
- Pseudomonas hormoni Sorty et al. 2023[22]
- Pseudomonas hygromyciniae Turner et al. 2023[23]
- Pseudomonas idahonensis Testerman et al. 2023[24]
- 印度假单胞菌 Pseudomonas indica Pandey et al. 2002
- 杰氏假单胞菌 Pseudomonas jessenii Verhille et al. 1999
- Pseudomonas jianii Yamada et al. 1968
- 晋州假单胞菌 Pseudomonas jinjuensis Kwon et al. 2003
- 基尔假单胞菌 Pseudomonas kilonensis Sikorski et al. 2001
- Pseudomonas knackmussii Stolz et al., 2007
- 韩国假单胞菌 Pseudomonas koreensis Kwon et al., 2003
- 昆明假单胞菌 Pseudomonas kunmingensis Xie et al. 2014
- Pseudomonas lalkuanensis Thorat et al. 2020[25]
- 亚麻假单胞菌 Pseudomonas lini Delorme et al., 2002
- Pseudomonas lutea Peix et al., 2004
- Pseudomonas marianensis Yang et al. 2023[26]
- 摩拉维亚假单胞菌 Pseudomonas moraviensis Tvrzová et al. 2006
- Pseudomonas nicosulfuronedens Li et al. 2021[27]
- 格陵兰假单胞菌 Pseudomonas nunensis Ntana et al. 2023[28]
- Pseudomonas oligotrophica Zhang et al. 2023[29]
- 耳炎假单胞菌 Pseudomonas otitidis Clark et al. 2006
- 海绵假单胞菌 Pseudomonas pachastrellae Romanenko et al. 2005
- 帕勒隆尼氏假单胞菌 Pseudomonas palleroniana Gardan et al. 2002
- Pseudomonas papaveris
- Pseudomonas paralcaligenes Ono et al. 2023[30]
- 海假单胞菌 Pseudomonas pelagia Hwang et al., 2009
- 烂泥假单胞菌 Pseudomonas peli Vanparys et al., 2006
- Pseudomonas perolens
- Pseudomonas petrae Nováková et al. 2023[31]
- 杀鱼假单胞菌 Pseudomonas piscis Liu et al. 2020
- 草假单胞菌 Pseudomonas poae Behrendt et al., 2003
- 食油假单胞菌 Pseudomonas pohangensis Weon et al., 2006
- Pseudomonas protegens Pseudomonas protegens
- 嗜冷假单胞菌 Pseudomonas psychrophila
- Pseudomonas psychrotolerans
- 魁北克假单胞菌 Pseudomonas quebecensis Tambong et al. 2023[32]
- Pseudomonas rathonis
- 食爬虫假单胞菌 Pseudomonas reptilivora
- Pseudomonas resiniphila
- Pseudomonas rhizosphaerae
- Pseudomonas rhodina Heumann 1962
- 浅红假单胞菌 Pseudomonas rubescens
- Pseudomonas rubra Testerman et al. 2023[33]
- 萨氏假单胞菌 Pseudomonas salomonii Gardan et al. 2002
- Pseudomonas schmalbachii Shelomi et al. 2021[34]
- Pseudomonas segitis
- 败血假单胞菌 Pseudomonas septica
- 塞尔维亚假单胞菌 Pseudomonas serbica Todorović et al. 2023[35]
- Pseudomonas serboccidentalis Todorović et al. 2023[35]
- 猴假单胞菌 Pseudomonas simiae Vela et al. 2006
- 猿假单胞菌 Pseudomonas solani Sawada et al. 2023[36]
- 松嫩平原假单胞菌 Pseudomonas songnenensis Zhang et al. 2015
- 猪假单胞菌 Pseudomonas suis
- Pseudomonas thermotolerans
- Pseudomonas toyotomiensis
- 山黄麻假单胞菌 Pseudomonas tremae
- 平凡假单胞菌 Pseudomonas trivialis
- Pseudomonas turbinellae
- Pseudomonas tuticorinensis
- Pseudomonas ullengensis Kim et al. 2021[21]
- 阴城假单胞菌 Pseudomonas umsongensis
- 温哥华假单胞菌 Pseudomonas vancouverensis
- 弗村假单胞菌 Pseudomonas vranovensis Tvrzová et al., 2006
- 黄色海假单胞菌 Pseudomonas xanthomarina Romanenko et al., 2005
- Pseudomonas xanthosomatis Girard et al. 2022,也拼作:Pseudomonas xanthosomae
- Pseudomonas xiamenensis Lai and Shao 2008
- 新疆假单胞菌 Pseudomonas xinjiangensis Liu et al. 2009
- 雄安假单胞菌 Pseudomonas xionganensis Zhao et al. 2020
- Pseudomonas yamanorum Arnau et al. 2015
- Pseudomonas yangonensis Tohya et al. 2020
- Pseudomonas zanjanensis Girard et al. 2022
- Pseudomonas zarinae Girard et al. 2022
- Pseudomonas zeae Girard et al. 2022
- Pseudomonas zeshuii Feng et al. 2012
- 肇东假单胞菌 Pseudomonas zhaodongensis Zhang et al., 2015
特征 编辑
假单胞菌属的总体特征如下:[37]
假单胞菌属的其他特征还有(部分种例外)在铁限制条件下分泌一种荧光铁载体。[38]有些假单胞菌属物种还可以分泌其他类型的铁载体,如绿脓杆菌可以分泌绿脓杆菌毒素[39],荧光假单胞菌可以分泌一种叫做thioquinolobactin的含硫铁载体。[40]假单胞菌属细菌氧化酶测试呈阳性,这也是其一个典型特征。
假单胞菌属可能是云中形成冰晶的最重要的成核剂,它们在全世界的雨雪形成过程中都起着十分重要的作用。[41]
生物膜形成 编辑
人们以前认为,假单胞菌属中所有的种都属于专性需氧微生物。近来人们在假单胞菌形成的生物膜中发现了例外。[42]在生物膜形成过程中,许多细胞都能够产生诸如藻朊酸盐等胞外多糖,这些多糖可以防止假单胞菌被白细胞吞噬。[43]胞外多糖还可以使假单胞菌在食物表面形成一片片难以去除的生物膜。腐坏食物表面的假单胞菌还会使食物闻起来有水果般的味道。
抗生素抗性 编辑
因为是革兰氏阴性细菌,大部分假单胞菌对青霉素以及很多相关β-内酰胺类抗生素都有抗性,但也有部分对哌拉西林、亚胺培南、替卡西林以及环丙沙星敏感。[43]临床治疗上的其他选择是诸如妥布霉素、庆大霉素以及阿米卡星等氨基糖苷类抗生素。
因为拥有含有孔蛋白的坚硬细胞壁,假单胞菌能够在严苛的环境下生存。它们的外排泵能够在许多抗生素發挥作用之前就把它们排出去。
绿脓杆菌是一种机会致病菌,对抗生素表现出低敏感性,较为棘手。[44]一是因为绿脓杆菌细胞壁渗透性低,二是因为其抗生素抗性基因(如mexAB-oprM、mexXY[45])可调控多种药物外排泵协同运作。 除了内生抗性,绿脓杆菌还很容易通过基因组突变或水平基因转移来产生新的抗性。若要获得对多种药物的抗性,就需要发生不同的突变或者接受多个水平基因的转移。高频突变倾向于在绿脓杆菌中产生能够引起慢性感染的抗生素抗性,而在内含子中同时出现多个抗生素抗性基因会使菌株同时具有多种抗性。有些研究发现,有些表型抗性会随生物膜形成或小的变种菌落而出现。[5]
对镓元素的敏感性 编辑
虽然镓元素没有什么自然的生物学功能,但它能对细胞过程产生类似于三价铁的影响。当细菌错误地将镓当成铁吸收后,镓并不像铁那样能够传递电子,这会影响细胞呼吸,最终导致细胞死亡。[46][47]
分类学 编辑
在二十世纪初,依靠经典分类方法来描述和鉴定这一微生物属经历了一段颇为坎坷的过程。当人们将比对核糖体RNA的大分子组分作为一项评判标准引入后,假单胞菌的分类就豁然开朗了:根据核糖体RNA的相似性,假单胞菌属可以分为五个核糖体RNA同源组。在Migula命名假单胞菌属数十年后,被划归到该属下的微生物种数目达到了惊人的比例。目前该属的微生物种类已经缩减到了不到原来的10%,这其中有不少菌是新命名的,而原来属于假单胞菌属的微生物,其实已经没有几个目前还在该属下了。依赖除核糖体RNA外的其他保守大分子的分类方法使得我们能够控制该属不至于过于庞大。[5]
致病性 编辑
动物病原体 编辑
致病菌包括绿脓杆菌、栖稻假单胞菌栖稻假单胞菌、和变形假单胞菌。绿脓杆菌是医院里一个较为头疼的问题,因为它对病人来说是侵染力排第二位的致病菌(院内感染)[來源請求]。它的致病机理可能和绿脓杆菌分泌的蛋白质有一定关系。该细菌具有种类非常广泛的分泌系统,它们分泌的许多蛋白质都和临床株的致病性有关。[48]
植物病原体 编辑
丁香假单胞菌是一类植物致病菌。它具有五十多种致病变种,其中许多具有高度的宿主专一性。假单胞菌属的许多其他物种也能成为植物致病菌,但是丁香假单胞菌是研究最为透彻的一个。
尽管并非严格的植物病原体,托拉斯假单胞菌在农业上也是一个头疼的问题,它可以造成栽培蘑菇的细菌性斑点病。[49]能够造成栽培蘑菇疾病的还有伞菌假单胞菌。[50]
作为生防试剂 编辑
二十世纪八十年代中期始,假单胞菌属中的某些物种就开始被用来抑制作物致病菌的生长或定殖。这种应用通常称作生物防治。荧光假单胞菌和Pseudomonas protegens的菌株(如CHAO或Pf-5)的生物防治特性是目前研究最为透彻的,尽管人们目前还不十分清楚荧光假单胞菌促进植物生长的过程是如何实现的。这其中可能的方式有:假单胞菌促进了宿主植物的系统抗性,使得植物能够更好地抵抗病原体;打败其他的土壤(致病)微生物(如分泌铁载体来获取铁元素);产生对其他土壤微生物有害的化合物,如吩嗪类抗体或氰化氢。上述所有方式均已得到了实验证实。[51]
其他具有生物防治功能的假单胞菌还有:绿叶假单胞菌,产生吩嗪类抗生素抑制特定真菌病原体生长[52];桔黄假单胞菌产生一种一直格兰仕阳性细菌的抗生素类物质di-2,4-diacetylfluoroglucylmethane。[53]
作为生物修复试剂 编辑
假单胞菌属的某些物种能够代谢环境污染物,它们可以用来进行生物修复。相关物种包括:
食物腐败 编辑
| 此章节需要扩充。 (2015年7月1日) |
因为具有多种多样的代谢特征,能够在低温和特殊环境下生存,很多假单胞菌都能够导致食物腐败。典型的例子有:莓实假单胞菌可以引起奶制品腐坏,[62],腐臭假单胞菌和霉味假单胞菌能够造成鸡蛋腐臭,[63]隆德假单胞菌,能够造成牛奶、奶酪、肉类和鱼类的腐败。[64]
以前属于该属的微生物种 编辑
最近,人们已经通过16s核糖体RNA测序将许多以前曾经属于假单胞菌属的微生物进行了重新分类。[6]这些被移出假单胞菌属的物种如下所示;点击相应物种原名会出现它们目前的新名字:
α 变形菌: P. abikonensis, P. aminovorans, P. azotocolligans, P. carboxydohydrogena, P. carboxidovorans, P. compransoris, P. diminuta, P. echinoides, P. extorquens, P. lindneri, P. mesophilica, P. paucimobilis, P. radiora, P. rhodos, P. riboflavina, P. rosea, P. vesicularis.
β 变形菌: P. acidovorans, P. alliicola, P. antimicrobica, P. avenae, P. butanovorae, P. caryophylli, P. cattleyae, P. cepacia, P. cocovenenans, P. delafieldii, P. facilis, P. flava, P. gladioli, P. glathei, P. glumae, P. huttiensis, P. indigofera, P. lanceolata, P. lemoignei, P. mallei, P. mephitica, P. mixta, P. palleronii, P. phenazinium, P. pickettii, P. plantarii, P. pseudoflava, P. pseudomallei, P. pyrrocinia, P. rubrilineans, P. rubrisubalbicans, P. saccharophila, P. solanacearum, P. spinosa, P. syzygii, P. taeniospiralis, P. terrigena, P. testosteroni.
γ-β 变形菌: P. beteli, P. boreopolis, P. cissicola, P. geniculata, P. hibiscicola, P. maltophilia, P. pictorum.
γ 变形菌: P. beijerinckii, P. diminuta, P. doudoroffii, P. elongata, P. flectens, P. halodurans, P. halophila, P. iners, P. marina, P. nautica, P. nigrifaciens, P. pavonacea,[65]P. piscicida, P. stanieri.
δ 变形菌: P. formicans.
噬菌体 编辑
能够侵染假单胞菌的噬菌体有很多,例如:
参考文献 编辑
- ^ Madigan M; Martinko J (编). Brock Biology of Microorganisms 11th. Prentice Hall. 2005. ISBN 0-13-144329-1.
- ^ Migula, W. (1894) Über ein neues System der Bakterien. Arb Bakteriol Inst Karlsruhe 1: 235–328.
- ^ Migula, W. (1900) System der Bakterien, Vol. 2. Jena, Germany: Gustav Fischer.
- ^ 4.0 4.1 4.2 Palleroni, N. J. The Pseudomonas Story. Environmental Microbiology. 2010, 12 (6): 1377–1383. PMID 20553550. doi:10.1111/j.1462-2920.2009.02041.x.
- ^ 5.0 5.1 5.2 5.3 Cornelis P (编). Pseudomonas: Genomics and Molecular Biology 1st. Caister Academic Press. 2008 [2016-12-13]. ISBN 1-904455-19-0. (原始内容存档于2016-09-12).
- ^ 6.0 6.1 Anzai Y; Kim H; Park, JY; Wakabayashi H. Phylogenetic affiliation of the pseudomonads based on 16S rRNA sequence. Int J Syst Evol Microbiol. 2000, 50 (4): 1563–89. PMID 10939664. doi:10.1099/00207713-50-4-1563.
- ^ Anzai, Y; Kudo, Y; Oyaizu, H. The phylogeny of the genera Chryseomonas, Flavimonas, and Pseudomonas supports synonymy of these three genera. Int J Syst Bacteriol. 1997, 47 (2): 249–251. PMID 9103607. doi:10.1099/00207713-47-2-249.
- ^ Yabuuchi, E.; Kosako, Y.; Oyaizu, H.; Yano, I.; Hotta, H.; Hashimoto, Y.; Ezaki, T.; Arakawa, M. Proposal of Burkholderia gen. Nov. And transfer of seven species of the genus Pseudomonas homology group II to the new genus, with the type species Burkholderia cepacia (Palleroni and Holmes 1981) comb. Nov. Microbiology and immunology. 1992, 36 (12): 1251–1275. PMID 1283774. doi:10.1111/j.1348-0421.1992.tb02129.x.
- ^ Yabuuchi, E.; Kosako, Y.; Yano, I.; Hotta, H.; Nishiuchi, Y. Transfer of two Burkholderia and an Alcaligenes species to Ralstonia gen. Nov.: Proposal of Ralstonia pickettii (Ralston, Palleroni and Doudoroff 1973) comb. Nov., Ralstonia solanacearum (Smith 1896) comb. Nov. And Ralstonia eutropha (Davis 1969) comb. Nov. Microbiology and immunology. 1995, 39 (11): 897–904. PMID 8657018. doi:10.1111/j.1348-0421.1995.tb03275.x.
- ^ Kim HS, Suh MK, Kim JS, Do HE, Eom MK, Jin JS, Lee JS. Pseudomonas aestuarii sp. nov., isolated from tidal flat sediment. Int J Syst Evol Microbiol 2023; 73:6190.
- ^ Testerman T, Varga J, Schiffer MM, Donohue H, Vieira Da Silva C, Graf J. Pseudomonas aphyarum sp. nov., Pseudomonas fontis sp. nov., Pseudomonas idahonensis sp. nov. and Pseudomonas rubra sp. nov., isolated from in, and around, a rainbow trout farm. Int J Syst Evol Microbiol 2023; 73:6201.
- ^ Banerjee S, Bedics A, Toth E, Kriszt B, Soares AR, Boka K, Tancsics A. Isolation of Pseudomonas aromaticivorans sp. nov from a hydrocarbon-contaminated groundwater capable of degrading benzene-, toluene-, m- and p-xylene under microaerobic conditions. Front Microbiol 2022; 13:929128.
- ^ Morimoto Y, Uwabe K, Tohya M, Hiramatsu K, Kirikae T, Baba T. Pseudomonas atagosis sp. nov., and Pseudomonas akappagea sp. nov., New Soil Bacteria Isolated from Samples on the Volcanic Island Izu Oshima, Tokyo. Curr Microbiol 2020; 77:1909-1915.
- ^ Zhang, D. C., Liu, H. C., Zhou, Y. G., Schinner, F., Margesin, R. (2011). Pseudomonas bauzanensis sp. nov., isolated from soil. Int J Syst Evol Microbiol 61 (Pt10): 2333-2337
- ^ Dong X, Rao Z, Wu S, Peng F, Xie Z, Long Y. Pseudomonas benzopyrenica sp. nov., isolated from soil, exhibiting high-efficiency degradation of benzo(a)pyrene. Int J Syst Evol Microbiol 2023; 73:6034.
- ^ Mohapatra B, Phale PS. Taxonomic, metabolic traits and species description of aromatic compound degrading Indian soil bacterium Pseudomonas bharatica CSV86T. Journal of Environmental Science and Health, Part A 2023; 58:633-646.
- ^ Liao K, Liu J, Gu YL, Wang C, Wei HL. Pseudomonas cucumis sp. nov., isolated from the rhizosphere of crop plants. Int J Syst Evol Microbiol 2023; 73:6208.
- ^ Mulet M, Martínez MJ, Gomila M, Dabernig-Heinz J, Wagner GE, et al. Genome-based species diversity assessment in the Pseudomonas chlororaphis phylogenetic subgroup and proposal of Pseudomonas danubii sp. nov. isolated from freshwaters, soil, and rhizosphere. Diversity 2023; 15:617.
- ^ Testerman T, Varga J, Schiffer MM, Donohue H, Vieira Da Silva C, Graf J. Pseudomonas aphyarum sp. nov., Pseudomonas fontis sp. nov., Pseudomonas idahonensis sp. nov. and Pseudomonas rubra sp. nov., isolated from in, and around, a rainbow trout farm. Int J Syst Evol Microbiol 2023; 73:6201.
- ^ Behrendt U, Ulrich A, Schumann P, Erler W, Burghardt J, Seyfarth W. A taxonomic study of bacteria isolated from grasses: a proposed new species Pseudomonas graminis sp. nov. Int J Syst Bacteriol 1999; 49:297-308.
- ^ 21.0 21.1 Kim CM, Jeong JW, Lee DH, Kim SB. Pseudomonas guryensis sp. nov. and Pseudomonas ullengensis sp. nov., isolated from soil. Int J Syst Evol Microbiol 2021; 71:5082.
- ^ Sorty AM, Zervas A, Garcia de Salamone IE, Nelson LM, Stougaard P. Pseudomonas hormoni sp. nov., a plant hormone producing bacterium isolated from Arctic grass, Ellesmere Island, Canada. Int J Syst Evol Microbiol 2023; 73:6119.
- ^ Turner TL, Mitra SD, Kochan TJ, Pincus NB, Lebrun-Corbin M, Cheung BH, Gatesy SW, Afzal T, Nozick SH, Ozer EA, et al. Taxonomic characterization of Pseudomonas hygromyciniae sp. nov., a novel species discovered from a commercially purchased antibiotic. Microbiol Spectr 2023; 11:e0183821.
- ^ Testerman T, Varga J, Schiffer MM, Donohue H, Vieira Da Silva C, Graf J. Pseudomonas aphyarum sp. nov., Pseudomonas fontis sp. nov., Pseudomonas idahonensis sp. nov. and Pseudomonas rubra sp. nov., isolated from in, and around, a rainbow trout farm. Int J Syst Evol Microbiol 2023; 73:6201.
- ^ Thorat V, Kirdat K, Tiwarekar B, DaCosta E, Debbarma P, Shouche Y, Sathe S, Goel R, Lodha T, Yadav A. Pseudomonas lalkuanensis sp. nov., isolated from a bacterial consortia of contaminated soil enriched for the remediation of e-waste. Int J Syst Evol Microbiol 2020; 70:6468-6475.
- ^ Yang Y, Gao Y, Liu Y, Liu B, Wang D, Xu Y, Wei Y. Pseudomonas marianensis sp. nov., a marine bacterium isolated from deep-sea sediments of the Mariana Trench. Arch Microbiol 2022; 204:638.
- ^ Li M, Ma Q, Kong D, Han X, Che J, Zhou Y, Jiang X, Ruan Z, Zhang Q. Pseudomonas nicosulfuronedens sp. nov., a nicosulfuron degrading bacterium, isolated from a microbial consortium. Int J Syst Evol Microbiol 2021; 71:4632.
- ^ Ntana F, Hennessy RC, Zervas A, Stougaard P. Pseudomonas nunensis sp. nov. isolated from a suppressive potato field in Greenland. Int J Syst Evol Microbiol 2023; 73:5700.
- ^ Zhang M, Li A, Yao Q, Xiao B, Zhu H. Pseudomonas oligotrophica sp. nov., a Novel Denitrifying Bacterium Possessing Nitrogen Removal Capability Under Low Carbon-Nitrogen Ratio Condition. Front Microbiol 2022; 13:882890.
- ^ Ono E, Tohya M, Watanabe S, Tada T, Kuwahara-Arai K, Oshiba A, Izumi N, Kirikae T. Pseudomonas paralcaligenes sp. nov., isolated from a hospitalized patient. Int J Syst Evol Microbiol 2023; 73:5649.
- ^ Novakova D, Koublova V, Sedlar K, Stankova E, Kralova S, Svec P, Neumann-Schaal M, Wolf J, Koudelkova S, Bartak M, et al. Pseudomonas petrae sp. nov. isolated from regolith samples in Antarctica. Syst Appl Microbiol 2023; 46:126424.
- ^ Tambong JT, Xu R, Chi SI, Birugu I, Bachelet S, Hutter C, Duceppe MO, Briere S. Pseudomonas quebecensis sp. nov., a bacterium isolated from root-zone soil of a native legume, Amphicarpaea bracteata (L.) Fernald, in Quebec, Canada. Int J Syst Evol Microbiol 2023; 73:5890.
- ^ Testerman T, Varga J, Schiffer MM, Donohue H, Vieira Da Silva C, Graf J. Pseudomonas aphyarum sp. nov., Pseudomonas fontis sp. nov., Pseudomonas idahonensis sp. nov. and Pseudomonas rubra sp. nov., isolated from in, and around, a rainbow trout farm. Int J Syst Evol Microbiol 2023; 73:6201.
- ^ Shelomi M, Chen WM, Chen HK, Lee HY, Young CC, Lin SY, Liaw SJ. Pseudomonas schmalbachii sp. nov., isolated from the gut of a millipede (Trigoniulus corallinus) from a coconut tree. Int J Syst Evol Microbiol 2021; 71:5101.
- ^ 35.0 35.1 Todorovic I, Abrouk D, Kyselkova M, Lavire C, Rey M, Raicevic V, Jovicic-Petrovic J, Moenne-Loccoz Y, Muller D. Two novel species isolated from wheat rhizospheres in Serbia: Pseudomonas serbica sp. nov. and Pseudomonas serboccidentalis sp. nov. Syst Appl Microbiol 2023; 46:126425.
- ^ Sawada H, Takeuchi K, Someya N, Morohoshi T, Satou M. Pseudomonas solani sp. nov. isolated from the rhizosphere of eggplant in Japan. Int J Syst Evol Microbiol 2023; 73:5942.
- ^ Krieg, Noel. Bergey's Manual of Systematic Bacteriology, Volume 1. Baltimore: Williams & Wilkins. 1984. ISBN 0-683-04108-8.
- ^ Meyer JM; Geoffroy VA; Baida N; Gardan, L.; et al. Siderophore typing, a powerful tool for the identification of fluorescent and nonfluorescent pseudomonads. Appl. Environ. Microbiol. 2002, 68 (6): 2745–2753. PMC 123936 . PMID 12039729. doi:10.1128/AEM.68.6.2745-2753.2002.
- ^ Lau GW; Hassett DJ; Ran H; Kong F. The role of pyocyanin in Pseudomonas aeruginosa infection. Trends in molecular medicine. 2004, 10 (12): 599–606. PMID 15567330. doi:10.1016/j.molmed.2004.10.002.
- ^ Matthijs S; Tehrani KA; Laus G; Jackson RW; et al. Thioquinolobactin, a Pseudomonas siderophore with antifungal and anti-Pythium activity. Environ. Microbiol. 2007, 9 (2): 425–434. PMID 17222140. doi:10.1111/j.1462-2920.2006.01154.x.
- ^ Biello, David (February 28, 2008) Do Microbes Make Snow? (页面存档备份,存于互联网档案馆) Scientific American
- ^ Hassett D; Cuppoletti J; Trapnell B; Lymar S; et al. Anaerobic metabolism and quorum sensing by Pseudomonas aeruginosa biofilms in chronically infected cystic fibrosis airways: rethinking antibiotic treatment strategies and drug targets. Adv Drug Deliv Rev. 2002, 54 (11): 1425–1443. PMID 12458153. doi:10.1016/S0169-409X(02)00152-7.
- ^ 43.0 43.1 Ryan KJ; Ray CG (编). Sherris Medical Microbiology 4th. McGraw Hill. 2004. ISBN 0-8385-8529-9.
- ^ Van Eldere J. Multicentre surveillance of Pseudomonas aeruginosa susceptibility patterns in nosocomial infections. J. Antimicrob. Chemother. February 2003, 51 (2): 347–352 [2016-12-13]. PMID 12562701. doi:10.1093/jac/dkg102. (原始内容存档于2009-02-10).
- ^ Poole K. Efflux-mediated multiresistance in Gram-negative bacteria. Clin. Microbiol. Infect. January 2004, 10 (1): 12–26 [2016-12-13]. PMID 14706082. doi:10.1111/j.1469-0691.2004.00763.x. (原始内容存档于2013-01-05).
- ^ "A Trojan-horse strategy selected to fight bacteria". INFOniac.com. 2007-03-16. Retrieved 2008-11-20.
- ^ Smith, Michael (2007-03-16). "Gallium May Have Antibiotic-Like Properties". MedPage Today. Retrieved 2008-11-20.
- ^ Hardie. The Secreted Proteins of Pseudomonas aeruginosa: Their Export Machineries, and How They Contribute to Pathogenesis. Bacterial Secreted Proteins: Secretory Mechanisms and Role in Pathogenesis. Caister Academic Press. 2009. ISBN 978-1-904455-42-4.
- ^ Brodey CL; Rainey PB; Tester M; Johnstone K. Bacterial blotch disease of the cultivated mushroom is caused by an ion channel forming lipodepsipeptide toxin. Molecular Plant–Microbe Interaction. 1991, 1 (4): 407–11. doi:10.1094/MPMI-4-407.
- ^ Young JM. Drippy gill: a bacterial disease of cultivated mushrooms caused by Pseudomonas agarici n. sp. NZ J Agric Res. 1970, 13 (4): 977–90. doi:10.1080/00288233.1970.10430530.
- ^ Haas D; Defago G. Biological control of soil-borne pathogens by fluorescent pseudomonads. Nature Reviews Microbiology. 2005, 3 (4): 307–319. PMID 15759041. doi:10.1038/nrmicro1129.
- ^ Chin-A-Woeng TF; Bloemberg, Guido V.; Mulders, Ine H. M.; Dekkers, Linda C.; et al. Root colonization by phenazine-1-carboxamide-producing bacterium Pseudomonas chlororaphis PCL1391 is essential for biocontrol of tomato foot and root rot. Mol Plant Microbe Interact. 2000, 13 (12): 1340–1345. PMID 11106026. doi:10.1094/MPMI.2000.13.12.1340.
- ^ Esipov; Adanin, VM; Baskunov, BP; Kiprianova, EA; et al. New antibiotically active fluoroglucide from Pseudomonas aurantiaca. Antibiotiki. 1975, 20 (12): 1077–81. PMID 1225181.
- ^ O'Mahony MM; Dobson AD; Barnes JD; Singleton I. The use of ozone in the remediation of polycyclic aromatic hydrocarbon contaminated soil. Chemosphere. 2006, 63 (2): 307–314. PMID 16153687. doi:10.1016/j.chemosphere.2005.07.018.
- ^ Yen KM; Karl MR; Blatt LM; Simon, MJ; et al. Cloning and characterization of a Pseudomonas mendocina KR1 gene cluster encoding toluene-4-monooxygenase. J. Bacteriol. 1991, 173 (17): 5315–27. PMC 208241 . PMID 1885512.
- ^ Huertas MJ; Luque-Almagro VM; Martínez-Luque M; Blasco, R.; et al. Cyanide metabolism of Pseudomonas pseudoalcaligenes CECT5344: role of siderophores. Biochem. Soc. Trans. 2006, 34 (Pt 1): 152–5. PMID 16417508. doi:10.1042/BST0340152.
- ^ Nojiri H; Maeda K; Sekiguchi H; Urata, Masaaki; et al. Organization and transcriptional characterization of catechol degradation genes involved in carbazole degradation by Pseudomonas resinovorans strain CA10. Biosci. Biotechnol. Biochem. 2002, 66 (4): 897–901. PMID 12036072. doi:10.1271/bbb.66.897.
- ^ Nam; Chang, YS; Hong, HB; Lee, YE. A novel catabolic activity of Pseudomonas veronii in biotransformation of pentachlorophenol. Applied Microbiology and Biotechnology. 2003, 62 (2–3): 284–290. PMID 12883877. doi:10.1007/s00253-003-1255-1.
- ^ Onaca; Kieninger, M; Engesser, KH; Altenbuchner, J. Degradation of alkyl methyl ketones by Pseudomonas veronii. Journal of Bacteriology. May 2007, 189 (10): 3759–3767. PMC 1913341 . PMID 17351032. doi:10.1128/JB.01279-06.
- ^ Marqués S; Ramos JL. Transcriptional control of the Pseudomonas putida TOL plasmid catabolic pathways. Mol. Microbiol. 1993, 9 (5): 923–929. PMID 7934920. doi:10.1111/j.1365-2958.1993.tb01222.x.
- ^ Sepulveda-Torres; Rajendran, N; Dybas, MJ; Criddle, CS. Generation and initial characterization of Pseudomonas stutzeri KC mutants with impaired ability to degrade carbon tetrachloride. Arch Microbiol. 1999, 171 (6): 424–429. PMID 10369898. doi:10.1007/s002030050729.
- ^ Pereira, JN & Morgan, ME. Nutrition and physiology of Pseudomonas fragi. J Bacteriol. Dec 1957, 74 (6): 710–3. PMC 289995 . PMID 13502296.
- ^ Levine, M & Anderson, DQ. Two New Species of Bacteria Causing Mustiness in Eggs. J Bacteriol. Apr 1932, 23 (4): 337–47. PMC 533329 . PMID 16559557.
- ^ Gennari, M & Dragotto, F. A study of the incidence of different fluorescent Pseudomonas species and biovars in the microflora of fresh and spoiled meat and fish, raw milk, cheese, soil and water. J Appl Bacteriol. Apr 1992, 72 (4): 281–8. PMID 1517169. doi:10.1111/j.1365-2672.1992.tb01836.x.
- ^ Van Landschoot, A.; Rossau, R.; De Ley, J. Intra- and Intergeneric Similarities of the Ribosomal Ribonucleic Acid Cistrons of Acinetobacter. International Journal of Systematic Bacteriology. 1986, 36 (2): 150. doi:10.1099/00207713-36-2-150.
- ^ 66.0 66.1 Hertveldt, K.; Lavigne, R.; Pleteneva, E.; Sernova, N.; Kurochkina, L.; Korchevskii, R.; Robben, J.; Mesyanzhinov, V.; Krylov, V. N.; Volckaert, G. Genome Comparison of Pseudomonas aeruginosa Large Phages (PDF). Journal of Molecular Biology. 2005, 354 (3): 536–545 [2016-12-13]. PMID 16256135. doi:10.1016/j.jmb.2005.08.075. (原始内容 (PDF)存档于2016-03-04).
- ^ Lavigne, R.; Noben, J. P.; Hertveldt, K.; Ceyssens, P. J.; Briers, Y.; Dumont, D.; Roucourt, B.; Krylov, V. N.; Mesyanzhinov, V. V.; Robben, J.; Volckaert, G. The structural proteome of Pseudomonas aeruginosa bacteriophage KMV. Microbiology. 2006, 152 (2): 529–534. PMID 16436440. doi:10.1099/mic.0.28431-0.
- ^ 68.0 68.1 Ceyssens, P. -J.; Lavigne, R.; Mattheus, W.; Chibeu, A.; Hertveldt, K.; Mast, J.; Robben, J.; Volckaert, G. Genomic Analysis of Pseudomonas aeruginosa Phages LKD16 and LKA1: Establishment of the KMV Subgroup within the T7 Supergroup. Journal of Bacteriology. 2006, 188 (19): 6924–6931. PMC 1595506 . PMID 16980495. doi:10.1128/JB.00831-06.
外部链接 编辑
| 维基共享资源上的相关多媒体资源:假单胞菌属 |
其他 编辑
- 假单胞菌对雨雪的形成作用
- 核反应堆中存活下来的假单胞菌(页面存档备份,存于互联网档案馆)
- 假单胞菌属基因组数据库 (页面存档备份,存于互联网档案馆)
- 假单胞菌属(页面存档备份,存于互联网档案馆)
- 荧光假单胞菌视频(页面存档备份,存于互联网档案馆)